Материал: Bovine Viral Diarrhea Virus Diagnosis, Management, and Control

66 |
BVDV: Diagnosis, Management, and Control |
sequence of viruses classified as BVDV led to the conclusion that there were actually two genotypes of BVDV, BVDV 1, and BVDV 2 (Pellerin et al., 1994; Ridpath et al., 1994a). The degree of sequence identity between pestiviruses in the 5’ UTR region is the most frequently used parameter for differentiation pestivirus species (Ridpath et al., 1994a; Harasawa and Mizusawa, 1995; Harpin et al., 1995; Harasawa, 1996; Baule et al., 1997; Becher et al., 1997; Giangaspero et al., 1997; Ridpath and Bolin, 1997; Sandvik et al., 1997; Vilcek et al., 1997; Wolfmeyer et al., 1997; Harasawa and Giangaspero, 1998; Ridpath and Bolin, 1998; Shimazaki et al., 1998; Letellier et al., 1999; Sakoda et al., 1999; Flores et al., 2000; Ridpath et al., 2000; Falcone et al., 2001; Tajima et al., 2001; Tajima et al., 2001; Vilcek et al., 2001b; Beer et al., 2002; Couvreur et al., 2002; Evermann and Ridpath, 2002; Flores et al., 2002). However, differences between BVDV 1 and BVDV 2 strains are found throughout the genome (Ridpath and Bolin, 1995b, 1997).
All pestiviruses are antigenically cross-reactive. Although antigenic differences do exist between pestivirus species, they are not extensive enough for pestivirus species to be recognized as serotypes. Convelescent sera generated against a virus from a particular pestivirus species will generally have a higher titer against other viruses from that species as opposed to viruses from the other pestivirus species (Ridpath, 2003). However, animal-to-animal variation and divergence among viruses from the same pestivirus species can make it difficult to reproducibly and reliably differentiate pestivirus species based on serology alone (Muller et al., 1997; Bolin and Ridpath, 1998). Mabs have been developed that differentiate between the pestivirus species (Wensvoort et al., 1989a; Zhou et al., 1989; Dahle et al., 1991; Edwards et al., 1991; Kosmidou et al., 1995). However, cross-reactivity between pestivirus species and variation within any one species make it difficult to generate a Mab that simultaneously differentiates between species and still recognizes all the viruses within one species. Antigenic variation is particularly pronounced among BVDV 1 and BVDV 2 strains (Ridpath et al., 1994a) and impacts on both detection and control.
In addition to the four recognized pestivirus species, three putative species have been suggested. These putative species are based on viruses isolated from a giraffe, a reindeer, and a pronghorn antelope (Becher et al., 1997; van Rijn et al., 1997; Harasawa et al., 2000; Vilcek et al., 2001b). Only one virus has been isolated for each of these putative species,
and no correlation with clinical disease has been reported.
THE PESTIVIRUS VIRION
Pestivirus virions are enveloped, spherical particles that are 40 to 60 nm in diameter. The virions are made up of a central capsid, composed of the virally encoded C protein and the genome RNA, surrounded by a lipid bilayer. The capsid appears as an electron-dense inner core with a diameter of approximately 30 nm (Horzinek et al., 1971). The structure and symmetry of the core has not been determined. Three virus encoded proteins—Erns, E1, and E2—are associated with the lipid bilayer envelope. The envelope surrounding the virion is pleomorphic, which impedes purification of infectious particles by banding in sucrose gradients and identification by electron microscopy. There appear to be 10–12 nm ring-like subunits on the surface of the virus envelope (Heinz et al., 2000). The Mr of the virion is estimated as 6.0 107, and the buoyant density in sucrose is 1.10–1.15 gm/cm3 (Heinz et al., 2000).
Virions are stable within a pH range of 5.7 to 9.3 (Hafez and Liess, 1972). Infectivity is not affected by freezing but decreases at temperatures above 40°C (Heinz et al., 2000). Like other enveloped viruses, BVDV are inactivated by organic solvents and detergents. Other methods of inactivation include Trypsin treatment (0.5 mg/ml, 37°C, 60 min) (Liess, 1990), ethylenimine (reduction of 5 log10 units using 10 mM at 37°C for 2 h) (Preuss et al., 1997), electron beam irradiation (4.9 and 2.5 kGy needed to reduce virus infectivity 1 log10 unit for frozen and liquid samples respectively) (Preuss et al., 1997), and gamma irradiation (20–30 kGy) (Miekka et al., 1998).
THE PESTIVIRUS GENOME
The pestivirus genome, in the absence of insertions, is approximately 12.3 Kb in length (Collett et al., 1988a, b; Moormann et al., 1990; Deng and Brock, 1992; Ridpath and Bolin, 1995b, 1997). The long open reading frame (approximately 4000 codons) is bracketed by relatively large 5’ (360–390 bases) and 3’ (200–240 bases) untranslated regions (UTR). The 5’ terminus does not contain a cap structure (Brock et al., 1992; Deng and Brock, 1993), and no poly(A) tract is present at the 3’ end. All pestivirus genomes terminate at the 3’ end with a short poly(C) tract. The highest nucleic acid sequence identity among pestiviruses is found in the 5’ UTR (Ridpath and Bolin,
Classification and Molecular Biology |
67 |
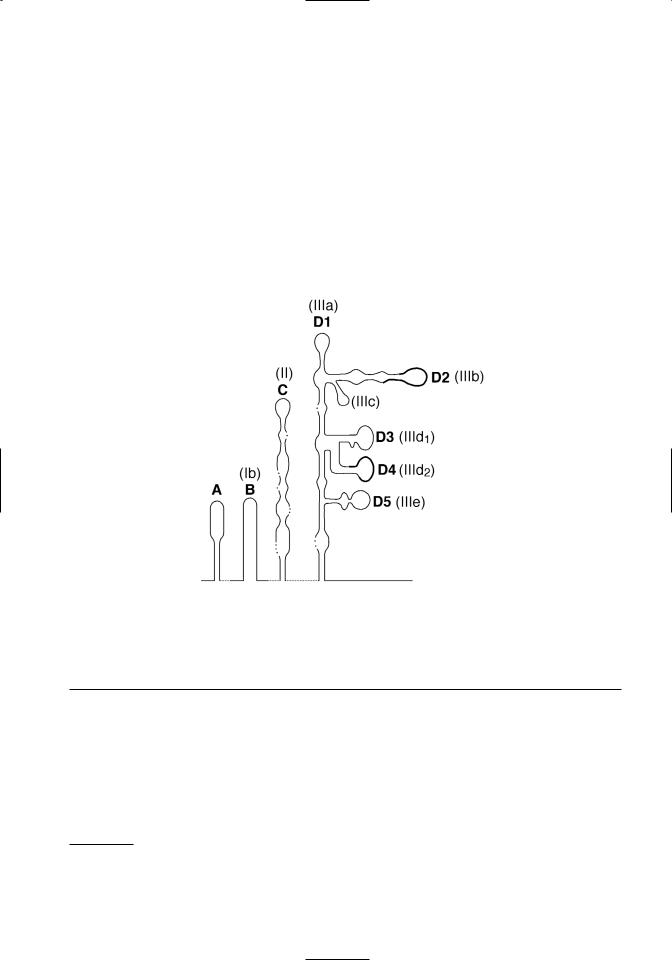
Figure 3.1. Conservation and predicted pseudoknots in pestivirus 5’ UTR sequences. Top. The alignment and comparison of 5’UTR sequences from recognized pestivirus species is shown. Hyperviable regions are boxed. Conserved sequences used in RT-PCR tests to detect pestiviruses are underlined. The first ATG of the open reading frame is in bold type. Bottom. A schematic of predicted tertiary structures in the 5’ UTR is shown. Deng and Brock (1993) domain designations are shown in bold, and Pestova and Hellen (1999) domain designations are shown in parentheses. Note that domain D5 of Deng and Brock is just a portion of the domain IIIe of Pestova and Hellen. The location of hypervariable regions shown in Figure 3.1a are denoted by a thickened line in the schematic.
1997). Although sequence identity is high among pestiviruses in the 5’ UTR, there are two short regions that are notable for their variability (Figure 3.1A). These are located between nucleotides 208– 223 and nucleotides 294–323 (nucleotide position numbers based on the sequence of BVDV 1-SD-11). Sequence variations in these regions have been ex-
ploited in PCR-based tests designed to differentiate BVDV genotypes (el-Kholy et al., 1998; Ridpath and Bolin, 1998). High conservation of 5’ UTR sequences is related to formation of tertiary structures required for internal ribosomal entry mediated initiation of translation (Deng and Brock, 1993; Pestova and Hellen, 1999) (Figure 3.1B).
1Although BVDV 1a-NADL and BVDV 2-890 are the type viruses for genotypes BVDV 1 and BVDV 2, respectively, both genomes have inserted sequences. Insertions can cause confusion when indicating genomic location based on nucleotide number. For this reason BVDV 1a-SD-1 is used as the reference for nucleotide position. It was the first noncytopathic BVDV1 sequenced and does not have an insertion. The accession number for BVDV 1a-SD-1 is M96751.

68 |
BVDV: Diagnosis, Management, and Control |
ENTRY OF THE VIRUS INTO CELLS, TRANSLATION, AND REPLICATION
The binding and entry of BVDV to susceptible cells has not been extensively studied. Based on the behavior of other flaviviruses and pestiviruses, it is hypothisized that binding and entry is a multistep process initiated by receptor-mediated endocytosis involving cell surface molecules and the viral proteins Erns and E2 (Schelp et al., 1995; Xue et al., 1997; Iqbal et al., 2000; Schelp et al., 2000; Iqbal and McCauley, 2002). The genomic RNA is uncoated following endocytosis and serves as the mRNA. No subgenomic mRNA molecules have been detected. Initiation of translation is mediated by a cap-independent internal initiation mechanism that requires an internal ribosome entry site (IRES) located within the 5’ UTR. A secondary structure model for the pestivirus 5’ UTR was first proposed by Deng and Brock (1993). Further refinements of a BVDV specific model have been developed (Poole et al., 1995; Pestova et al., 1998; Pestova and Hellen, 1999). Unfortunately a universal terminology system was not adopted. The different names used for the same structural domains by different research groups can be confusing. The proposed 5’ UTR secondary structures shown in Figure 3.1B are identified by both the domain names given by Deng and Brock and those given by Pestova and Hellen. It should be noted that domain D5 of Deng and Brock is just a portion of the IIIe domain of Pestova and Hellen. The IIIe domain contains the D5 domain plus the nucleotides between D5 and nucletide 380 (numbering based on BVDV 1a-SD-1). Domains C (II) and D (III) contain structures critical for the attachment of the initiation complex. A pseudoknot formed just upstream of the start codon is necessary for IRES function. The hypervariable regions used in PCR based tests to differentiate BVDV genotypes are located in domains D2 (IIIb) and D4 (IIId2).
VIRAL PROTEINS
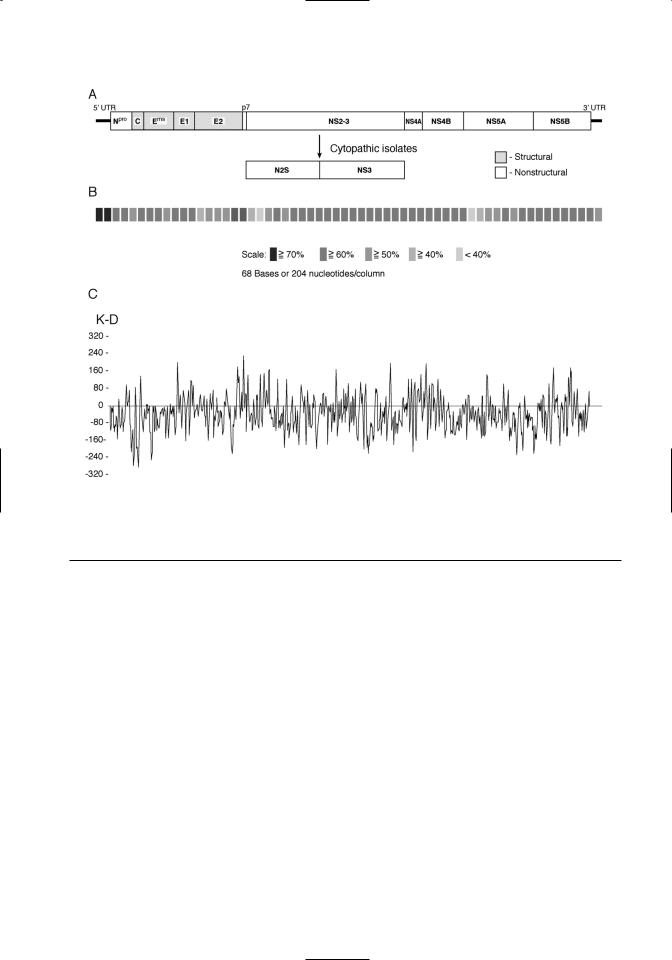
The large open reading frame is translated as a polyprotein. The order of the individual viral proteins within the polyprotein is as follows: Npro- C-Erns-E1-E2-p7-NS2/3-NS4A-NS4B-NS5A- NS5B (Figure 3.2A). The polyprotein is processed cotranslationally and posttranslationally by host and viral proteases. The size, function, and characteristics of the viral proteins are summarized in Table 3.1. Conservation of the polyprotein coding sequences among pestiviruses is shown in Figure 3.2B. The predicted hydrophobicity profile of the
polyprotein is shown in Figure 3.2C. The hydrophobicity profile of the polyprotein is conserved within the pestivirus genus.
Viral structural proteins
The proteins associated with the mature virion are C, Erns, E1, and E2. C is the virion nucleocapsid protein. It is highly basic and relatively conserved among the pestivirus species. It is postulated that the C terminus of the nucleocapsid protein contains an internal signal sequence that directs translocation of structural glycoproteins to the endoplasmic reticulum (Heinz et al., 2000). The Erns protein is glycosylated and forms disulfide-linked homodimers (Weiland et al., 1990). In tissue culture systems Erns may be found both associated with released virus and free in the culture medium. This protein has an unusual ribonuclease activity. Although the function of this ribonuclease activity in the viral life cycle is unknown, antibodies that inhibit ribonuclease activity neutralize virus infectivity of classical swine fever viruses (CSFV) (Windisch et al., 1996). The ribonuclease activity does not require dimer formation or glycosylation (Windisch et al., 1996). The Erns of CSFV possesses epitopes that induce neutralizing antibodies (hereafter referred to as neutralizing epitopes) and CSFV Erns subunit vaccines induce neutralizing antibodies (Konig et al., 1995). It is not known whether the Erns of BVDV possesses neutralizing epitopes that are important in disease control. However, protection afforded by killed BVDV vaccines does not appear to be dependent upon Erns specific antibodies (Bolin and Ridpath, 1990).
Both the E1 and E2 proteins possess potential membrane-spanning domains and are glycosylated. They are predicted to be integral membrane proteins and interact to form heterodimers (Weiland et al., 1990). The E2 protein is the immunodominant structural protein and possesses neutralizing epitopes (Donis et al., 1988; Bolin and Ridpath, 1989). Mabs produced against the E2 have been used to differentiate pestivirus species (Peters et al., 1986; Wensvoort et al., 1986; Bolin et al., 1988; Hess et al., 1988; Wensvoort et al., 1989a; Corapi et al., 1990a; Edwards and Sands, 1990; Dahle et al., 1991; Deregt et al., 1994; Paton et al., 1994; Ridpath et al., 1994a; Ridpath and Bolin, 1997). The coding region for E2 neutralizing epitopes are found in a hypervariable region located in the N-terminal portion of the E2 protein (Paton et al., 1992; Deregt et al., 1998) (refer to Figure 3.2B). E2 antigenic variation is particularly pronounced among BVDV strains and contributes to vaccine failure (Bolin et al., 1988; Bolin and Rid-
Classification and Molecular Biology |
69 |
Figure 3.2. The pestivirus genome. A. Organization of the pestivirus genome is shown. Structural proteins are denoted by shading. B. Sequence conservation among pestivirus species is shown. These results are a composite of comparisons between the four recognized pestivirus species. Sequence similarity is indicated by different levels of shading. C. A hydrophobicity plot of a pestivirus consensus sequence is shown. A consensus sequence was derived by comparison of representatives of noncytopathic strains from each of the four recognized pestivirus species.
path, 1989, 1990; Bolin et al., 1991a; Bolin et al., 1994; Ridpath et al., 1994a; Van Campen et al., 1997; Bolin and Ridpath, 1998; Van Campen et al., 1998; Ridpath et al., 2000; Van Campen et al., 2000).
Viral nonstructural proteins
The first viral protein encoded by the BVDV ORF is the nonstructural protein, Npro. This protein, which is unique to the pestivirus genus, is an autoprotease whose only known function is to cleave itself from the polyprotein. The next nonstructural protein, p7, follows the structural protein E2 in the polyprotein (Elbers et al., 1996). It consists of a central charged region flanked by hydrophobic termini. The role of this cell-associated protein is unknown. It is hypothesized that p7 is required for production of infectious virus but not for RNA replication (Harada et al., 2000). It is inefficiently cleaved from the E2, leading to two intracellular forms of E2 with different C termini (E2 and E2-p7) (Elbers et al., 1996).
Neither p7 nor E2-p7 are found associated with infectious virus.
Following p7 the next nonstructural protein in the polyprotein is the serine protease, NS2-3. In BVDV strains from the cytopathic biotype (see discussion of biotype below), the NS2-3 is cleaved to NS2 and NS3. The serine protease activity of the NS2-3 resides in the NS3 portion of the protein. The function of the NS2 is unknown. It is not required for RNA replication, and its cleavage from the NS2-3 does not affect serine protease activity (Behrens et al., 1998). Sequence analysis reveals regions in the NS2 with homology to zinc-finger motifs present in DNA binding proteins (De Moerlooze et al., 1991). However, a function for the putative DNA binding activity of NS2 in viral replication has not been demonstrated.
Both the uncleaved NS2-3 and the cleaved NS3 act as serine proteases (Tautz et al., 1997) that cleave the remaining nonstructural proteins from the

70 |
|
BVDV: Diagnosis, Management, and Control |
||
Table 3.1. Pestivirus proteins |
|
|
||
|
Estimated |
|
|
|
Viral |
Size |
|
Neutralizing |
|
Protein |
(K Daltons) |
Attributes |
Epitope(s) |
Function |
|
|
|
|
|
Npro |
020 |
Nonstructural |
N |
Autoproteolysis |
|
|
|
|
Not required for RNA replication |
C |
014 |
Structural |
N |
Forms nucleocapsid of virion |
|
|
Conserved |
|
|
Erns |
|
Highly basic |
|
|
048 |
Structural |
Y |
Envelope-associated glycoprotein |
|
|
|
7–9 glycosylation sites |
|
Ribonuclease activity |
E1 |
025 |
Structural |
N |
Envelope-associated glycoprotein |
|
|
2–3 glycosylation sites |
|
Integral membrane protein |
E2 |
053 |
Structural |
Y |
Envelope-associated glycoprotein |
|
|
4–6 glycosylation sites |
|
Integral membrane protein |
|
|
Least conserved of structural proteins |
Immunodominant structural protein |
|
p7 |
007 |
Nonstructural |
N |
Function unknown |
|
|
Central charged region flanked |
|
Required for production of |
|
|
by hydrophobic termini |
|
infectious virus but not RNA |
|
|
|
|
replication |
NS 2/3 |
125 |
Nonstructural |
N |
NS2 has a zinc-finger–like domain |
NS2 |
054 |
In cytopathic biotype NS 2/3 cleaved |
NS2/3 and NS3 contain RNA |
|
NS3 |
080 |
to NS2 and NS3 |
|
helicase and N-terminal serine |
|
|
Conserved |
|
protease domains; cleaves itself |
|
|
|
|
and remaining nonstructural pro- |
|
|
|
|
teins from viral polyprotein |
|
|
|
|
Immunodominant nonstructural |
|
|
|
|
protein |
NS4A |
7.2 |
Nonstructural |
N |
Serine protease cofactor |
|
|
Hydrophobic |
|
|
NS4B |
38–39 |
Nonstructural |
N |
Replicase component |
|
|
Hydrophobic |
|
|
NS5A |
55–56 |
Nonstructural |
N |
Replicase component |
|
|
Phosphorylated |
|
|
NS5B |
81–82 |
Nonstructural |
N |
RNA-dependent RNA polymerase |
polyprotein. Purified BVDV NS3 also possesses RNA helicase and RNA-stimulated NTPase activities (Tamura et al., 1993; Warrener and Collett, 1995). All three activities (serine protease, RNA helicase, and RNA-stimulated NTPase) are essential to virus viability (Grassmann et al., 1999; Gu et al., 2000). Antibodies to the NS2-3 and NS3 do not neutralize infectivity. However, the NS2-3 and NS3 (but not the NS2) are strongly recognized by polyclonal convalescent sera (Donis et al., 1991). Animals vaccinated with modified live vaccines also have a strong antibody response to the NS2-3 and/or NS3 protein (Bolin and Ridpath, 1989). In contrast, animals vaccinated with inactivated (killed) vaccines
primarily react with structural proteins and not the NS2-3 or NS3 (Bolin and Ridpath, 1990). The difference in recognition of NS2-3 or NS3 may be useful in differentiating between immune responses to inactivated vaccines and immune responses to natural infection.
The NS4A and NS4B proteins are similar in size, composition and hydrophobicity to the NS4A and NS4B proteins of other flaviviruses (Lindenbach and Rice, 2001). NS4A acts as a cofactor for the NS2-3 and NS3 serine protease activity. NS4B and NS5A probably are replicase complex components. RNA polymerase activity has been demonstrated for the NS5B protein (Lai et al., 1999).