Материал: Bovine Viral Diarrhea Virus Diagnosis, Management, and Control

180 |
BVDV: Diagnosis, Management, and Control |
Taken together, these results suggest that attachment and entry of BVDV into a cell is a multistep process. First, the virus attaches to the cell surface through interaction of Erns envelope protein and a docking glycosaminoglycan receptor molecule. This step can occur in both susceptible and nonsusceptible cells because these molecules are present on most cells. It appears that binding to the docking receptor is insufficient by itself to bring about uptake of the virus (Flores and Donis, 1995; Flores et al., 1996). Entry into the cell is further mediated by attachment of the E2 envelope protein to the LDLR receptor and internalization via endocytosis. This is probably the rate-limiting step due to the relatively low abundance of the LDLR on most cell types (Agnello et al., 1999). As was stated earlier, the presence of other, minor receptor molecules cannot be ruled out at this time.
INTERACTIONS WITH CELLULAR FACTORS DURING REPLICATION
Upon entry into the cell, the genomic RNA must act as mRNA, directing the translation of viral proteins. Viral proteins participate in the necessary functions for RNA replication, protein processing (protease cleavages), and protein trafficking, but are insufficient by themselves to bring about all of these events. Recruitment of cellular factors is necessary for the successful completion of the replication process. Many of these factors are known, but the identity and function of others remain elusive.
Probably the most important of the cellular functions utilized by BVDV for replication is protein synthesis. This process is much too complex to be encoded by a simple virus. Thus, BVDV must utilize the existing translational factors employed by the cell. In noninfected cells, eukaryotic capdependent translation initiation begins by the binding of eIF4E (the cap binding protein), as part of the heterotrimeric eIF4F complex (composed of eIF4A, eIF4E, and eIF4G), to the 5’ m7G cap of the mRNA molecule (Figure 11.1A). The 40S small ribosome subunit, associated with the ternary complex eIF2/ GTP/Met-tRNAi, eIF2 and eIF3, forms the 43S preinitiation complex. The binding of the 43S subunit to the mRNA is directed by the eIF4F complex and results in the formation of the 48S complex. The ATP-dependent helicase activity of the eIF4A subunit is thought to unwind the RNA, allowing the binding of the 43S complex to the mRNA.
Both eIF1 and eIF1a are necessary for proper assembly of the 48S preinitiation complex as well as
Figure 11.1. Translation initiation of cellular transcripts as compared to that of the pestivirus genomic RNA. A. Eukaryotic translation initiation takes place by binding of the cap structure (M7G) as directed by the eIF4F complex. The attachment of the 40S small ribosomal subunit and other factors necessary to form the 48S preinitiation complex are illustrated. The scanning competent complex moves down the transcript until encountering the initiation codon, at which point eIF5 and eIF5B direct formation of the 80S ribosomal complex and initiation of translation. B. Formation of the 48S preinitiation complex at the BVDV IRES requires only the 40S small ribosomal subunit, eIF3 and eIF2/GTP/ Met-tRNAi. This results in the assembly of the transla- tion-competent 80S ribosomal complex at the initiator AUG and translation of the BVDV polyprotein.
making the 48S preinitiation complex competent for scanning to the initiation codon. The scanning process to the initiation codon also requires eIF4B and eIF4F. At the initiation codon, eIF5, through interaction with both eIF2 and eIF3, causes the hydrolysis of the GTP moiety of eIF2, resulting in the
Interactions of Virus and Host |
181 |
release of eIF2-GDP. Initiation factor eIF5B, again with hydrolysis of GTP, directs the attachment of the 60S ribosomal subunit in the formation of the translationally competent 80S ribosome. For a more detailed discussion of the eukaryotic protein translation initiation process, see Pestova et al. (1998) and Pestova et al. (2001).
POLYPROTEIN TRANSLATION
Following the uptake and the release of the viral genome, the next critical event in the replication cycle is the production of viral proteins, both those necessary for replication and those that interact with host factors to generate a cellular environment optimized for production of virus progeny. The BVDV genomic RNA functions as mRNA for the immediate synthesis of a large viral polyprotein that is encoded in the single open reading frame (ORF) of the virus. The ORF encodes a large protein of approximately 4000 amino acids that is coand posttranslationally cleaved to produce the individual, mature proteins. The proteins, in order of placement within the genome from N- to C-terminus are NH2-Npro- capsid-Erns-E1-E2-p7-NS2/3-NS4a-NS4b-NS5a- NS5b-COOH. The envelope glycoproteins—Erns, E1, and E2—are translocated into the lumen of the endoplasmic reticulum for further posttranslational modification (Rümenapf et al., 1993). The E1 and E2 are inserted into the membrane of the endoplasmic reticulum during translocation while Erns remains soluble.
A major difference between the BVDV genomic RNA and most cellular mRNA molecules is the presence of a long 5’ nontranslated region (5’ UTR) containing a high degree of secondary structure (Figure 11.2) as well as multiple upstream AUG codons. This roughly 390 base region, termed the internal ribosome entry site (IRES), functions to direct the attachment and assembly of the ribosome at the initiation codon of the ORF, insuring that initiation of translation begins at the proper AUG codon.
Computational analysis of the 5’ untranslated region of pestivirus genomes revealed the presence of higher order folding and secondary structure (Brown et al., 1992; Deng and Brock, 1993; Le et al., 1995). The presence of multiple AUG codons was noted, several with stronger matches to the Kozak consensus sequence (Kozak, 1987) than the authentic initiation codon (Deng and Brock, 1993). These analyses resulted in the development of a model of secondary structure that revealed a series of stem-loops that were highly conserved structurally between BVDV and CSFV. These tertiary structures were conserved despite the sequence divergence between the two viruses. Brown et al. (1992) used doubleand single-strand specific RNases to confirm the structure of the stem-loop structures and further found them to be highly similar to that described for the IRES of HCV (Tsukiyama-Kohara et al., 1992). Modeling studies by Le et al. (1995) predicted stem-loop structures as well as a psuedoknot in the BVDV and CSFV 5’
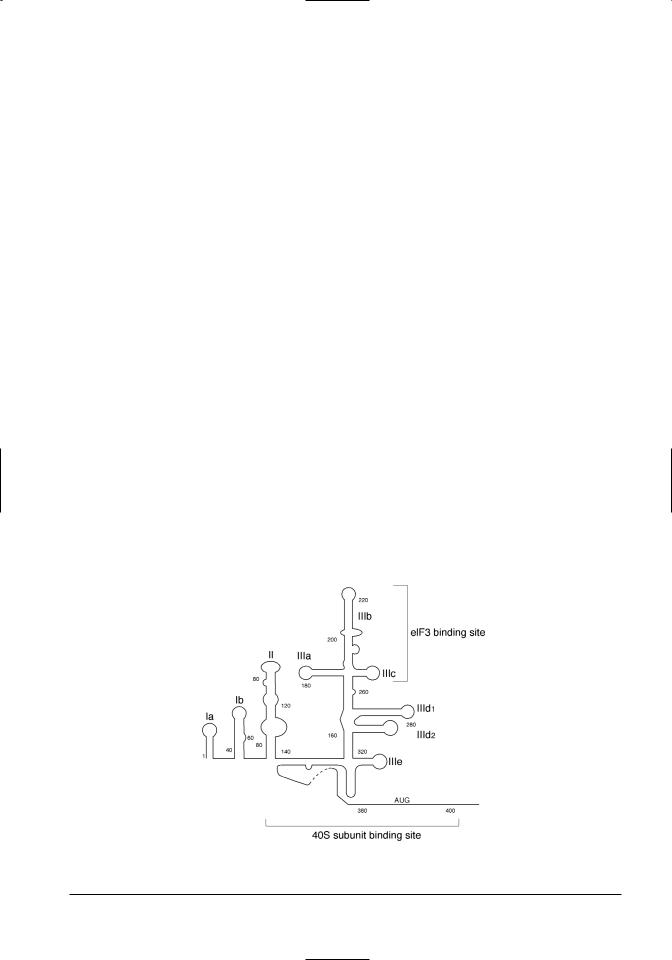
Figure 11.2. The secondary structure of the 5’ UTR/IRES of the BVDV genomic RNA. The conserved stem-loop structures of this region are illustrated, along with the location of the binding sites of eIF3 and the 40S ribosomal subunits (Pestova et al., 2001). The nucleotide sequence is not shown because of variation between BVDV strains.

182 |
BVDV: Diagnosis, Management, and Control |
UTRs that were consistent with prior RNase sensitivity studies (Brown et al., 1992). The pestivirus IRES also contained a structural motif that was proposed to base pair with the 3’ end of the 18S rRNA, in a manner similar to that described for the picornaviruses (Deng and Brock, 1993; Le et al., 1995). This base pairing would occur at sequences immediately 5’ of the AUG initiation codon. It was proposed that this base pairing between the viral and 18s rRNA functioned in initiation of cap-independent translation by positioning of the ribosome.
The first description of the 5’ UTR as a functional IRES was by Poole et al. (1995). In vitro transcription of the 5’ UTR along with the p20 (Npro) coding sequences of the BVDV ORF yielded expression of the p20 protein in an in vitro cell-free expression system. Inclusion of the 5’ UTR as a spacer between the CAT and luciferase genes in a bicistronic construct yielded expression of both genes. Expression studies using constructs with both genes without a spacer resulted in only CAT expression. Deletion of nucleotides 173 through 236 of the 5’ UTR reduced expression of luciferase by about two-thirds. These results suggested that the BVDV 5’ UTR possesses IRES activity similar to that of HCV. Rijnbrand et al. (1997), in a separate study, demonstrated that the CSFV IRES sequences functioned similarly when used as a spacer in a bicistronic construct. Mutational analysis of the nucleotide sequences that formed the pseudoknot revealed that RNAs unable to form base pairs in stem II of the pseudoknot were translationally inactive. However, translational activity could be restored by introduction of compensatory base changes that restored the stem loop structure (Rijnbrand et al., 1997; Fletcher and Jackson, 2002). Neither an AUG codon 7 base upstream of the authentic initiation AUG codon nor an AUG downstream could initiate translation. This indicates that scanning by the ribosome on the IRES is limited to a very small region.
A genetic approach to dissection of BVDV IRES function (Chon et al., 1998) using mutational analysis through a number of in vitro–introduced deletions or insertions, showed that stem loops Ia and Ib were dispensable for efficient translation, and IIIb and IIIe were partially required (Figure 11.3). Deletions or insertions in II, IIIa, IIIc or IIId could cause a tenfold or greater reduction in translation. Confirmation of these data was provided by chemical and enzymatic footprinting (Sizova et al., 1998), which demonstrated that the translation initiation factor eIF3 bound to and protected distinct regions of domain III, particularly IIIb and IIIc (refer to
Figure 11.2). Further, deletions of these sequences abrogated translation. They suggested that eIF3 bound the IRES in a sequenceor structure-specific manner. In UV cross-linking experiments to determine which subunits of the 10 protein subunit eIF3 complex actually contacts the IRES, 4 subunits were found to be labeled by the radiolabeled RNA (p170, p116, p66, and p47). The p116 and p66 subunits both contain RNA recognition motifs and thus are likely to be determinants of the interaction of eIF3 with the IRES (Sizova et al., 1998).
Pestova and Hellen (1999) provided evidence that the IRES was bound independently of eIF3 and the 40S ribosomal subunit. The IRES contains complex structure determinants (refer to Figure 11.2) that mediate attachment of the 48S ribosomal subunit complexes to the initiation codon (Pestova et al., 1998). Determinants of domain III mediate eIF3 binding. Pestivirus translation initiation is dependent on eIF3, eIF2, GTP, and Met-tRNAmet (48S subunit assembly) but not on any constituent of the eIF4G subunit (refer to Figure 11.1B). eIF3 enhances 48S complex formation, probably by stabilizing ribosomal complexes by its interaction with the IRES, and is absolutely required for subsequent subunit joining to form active 80S ribosomes (Sizova et al., 1998). The probable function of eIF3 is to recruit factors eIF1 and eIF5 to the 48S complex (Pestova and Hellen, 1999). Using cryoelectron microscopy, Spahn et al. (2001) demonstrated that binding of the HCV IRES caused a conformational change in the 40S ribosomal subunit that resulted in closing of the mRNA binding cleft, and in IRESmediated positioning of the initiation codon in the P site of the ribosome. This was the first description of a viral RNA that actively manipulates the structure of the transcriptional machinery, resulting in the promotion of translation initiation without the assistance of otherwise necessary initiation factors (Spahn et al., 2001).
Several groups have reported that initiation factors eIF4A, eIF4B, and eIF4F were not required for translation initiation from the pestivirus IRES (Rijnbrand et al., 1997; Chon et al., 1998; Pestova and Hellen, 1999; Sizova et al., 1998; Fletcher et al., 2002). These findings were derived from different types of experiments, including reconstituted in vitro translation initiation systems (Pestova and Hellen, 1999) and coexpression of picornavirus 2A protease (Rijnbrand et al., 1997; Chon et al., 1998). In addition, this IRES-driven translation was shown to be independent of the translation initiation factor eIF-4F by coexpression of the poliovirus 2A protease

Interactions of Virus and Host |
183 |
that cleaves eIF4F. This protease-inhibited capdependent translation in picornavirus-infected cells (Rijnbrand et al., 1997; Chon et al., 1998; Pestova et al., 1998; Pestova et al., 2001; Fletcher et al., 2002). The report by Pestova et al. (1998) was the first describing the lack of reliance by the pestivirus IRES on canonical translation initiation factors that are necessary for normal cellular cap-dependent translation initiation. It was concluded that pestivirus protein translation initiation is simpler than that employed by the picornaviruses and more closely resembles prokaryotic translation initiation. Translation initiation from the flavivirus or pestivirus IRES requires so few translation initiation factors that a simpler, equally efficient means of translation initiation is unlikely (Pestova et al., 1998). Cellular RNAs that contain an IRES are generally those that are expressed under stress conditions (i.e., heat shock, reactive oxygen species) when the activities of eIF4B, E, and F are reduced (Pestova et al., 1998).
The 5’ end of the IRES was mapped to between nucleotides 28 and 75 (Rijnbrand et al., 1997; Chon et al., 1998). The 3’ end of IRES extends into Npro coding sequences of the ORF (Chon et al., 1998) as far as 51 bases (Fletcher et al., 2002) and possesses single-strandedness that appears important for ribosome binding and translation initiation. Studies mapping the termini of the IRES were done utilizing in vitro systems where the IRES was directly fused to the ORF encoding the expression reporter. Additional work showed that efficient IRES function, in the context of translation from a functional replicon, required the 5’ Ia stem-loop structure (Yu et al., 2000). Utilizing the subgenomic RNA replicon DI9c (Behrens et al., 1998), translation was shown to be inefficient when the Ia stem-loop structure was mutated or missing. In addition, it was found that the Ia structure was important in RNA replication as well. The authors postulated that this structural motif may be involved in RNA:RNA interactions that may be necessary for committing the RNA molecule to function as template for either translation or RNA replication.
PROCESSING OF ENVELOPE GLYCOPROTEINS
BY CELLULAR ENZYMES
The BVDV polyprotein, encoded by the single ORF of the genomic RNA, is co-and posttranslationally processed by both host and viral proteases to yield the mature viral proteins. The protease cleavages of the nonstructural proteins are carried out by the NS2/3 serine protease (Wiskerchen and Collett, 1991; Tautz et al., 1997; Xu et al., 1997). However,
the processing of the virus structural proteins, encoded in the N-terminal one-third of the polyprotein, is carried out by host signal peptidases (Figure 11.3A) following translocation into the lumen of the endoplasmic reticulum (Figure 11.3; Rümenapf et al., 1993; Lin et al., 1994). A model of this translocation and proteolytic processing has been proposed (Rümenapf et al., 1993). Cotranslationally, the envelope proteins are translocated into the ER utilizing a signal sequence located between the capsid and Erns proteins. In doing so, the ribosome attaches to the Sec61p complex, a protein translocation channel in the ER membrane. The signal peptide is inserted into the membrane and the nascent protein chain is translocated into the lumen of the ER in a loop structure (Matlack et al., 1998). A rapid signalase cleavage occurs at the N-terminus of the Erns protein, possibly leaving the capsid protein attached to the cytoplasmic side of the membrane.
The core (capsid) protein of HCV is released from the endoplasmic reticulum membrane by the cellular protease SPP (McLauchlan et al., 2002), and it is likely that process occurs in a similar manner in BVDV-infected cells. Transfer of the remaining Erns/E1/E2 envelope protein precursor continues with interruption by two hydrophobic domains at the end of the E1 protein. The first hydrophobic domain acts as a stop transfer for the nascent protein, and the second acts as a signal sequence to begin translocation of the E2 protein. Both may function to anchor the E1 protein into the membrane. There are similar stop/start transfer signals at the C- terminus of the E2 region. The internal signal sequences of the Erns/E1/E2 precursor are the only determinants for compartmentalization of the mature envelope proteins (Wu, 2001). Cleavage of the Erns/E1/E2 precursor from the downstream translation product occurs rapidly. Processing of the precursor begins with the signalase cleavage at the E1/E2 border. This is followed by the separation of the Erns/E1 proteins and is probably dependent on a conformational change taking place following the initial E2 cleavage reaction. The Erns protein forms homodimers by intermolecular disulfide bond formation and, having no hydrophobic transmembrane domain, remains soluble and associates with the E1E2 heterodimers by an unknown mechanism (Hulst and Moorman, 2001). Unassociated Erns is also secreted into the extracellular domain and can be found free in the medium of infected cells cultured in vitro (Rümenapf et al., 1993). As discussed previously, Erns is essential for virus attachment and uptake (Hulst and Moormann, 1997).

184 |
BVDV: Diagnosis, Management, and Control |
Figure 11.3. Proposed mechanism for transporting and processing of pestivirus envelope proteins. A. Genomic organization of the BVDV genome showing cleavage sites of the mature proteins with those cleaved by host signalases denoted by asterisks. The sizes of the proteins are not drawn to scale. B. Import into the lumen of the endoplasmic reticulum and cleavage of the mature proteins from the polyproteins result in the release of free Erns into the lumen of the endoplasmic reticulum, while cleavage of the E1 and E2 proteins results in retention of the membrane anchors in the membrane, probably facilitating downstream processing, heterodimerization and transport.
The E1 and E2 proteins form disulfide-linked heterodimers after proper folding of the proteins in association with the endoplasmic reticulum protein calnexin (Branza-Nichita et al., 2001). The folding of these proteins is dependent on intramolecular disulfide bridge formation and varies in the rate this occurs between the two envelope proteins. The folding of the E2 protein is rapid, showing completion within 2.5 minutes; the folding of the E1 proteins takes approximately 30 minutes. Thus the rate of folding of the E1 protein is the rate-limiting step in the formation of E1-E2 dimers. The interaction of envelope proteins with the lectin-like calnexin molecule is dependent on the removal of glucose moieties on the N-linked glycans by endoplasmic reticulum -glucosidases (Branza-Nichita et al., 2001). If the removal of the glucose moieties does not occur, interaction with calnexin is inhibited and the improperly folded proteins are targeted for degradation by the proteosome (Branza-Nichita et al., 2002). Decrease in size of cleaved proteins that indicated trimming of sugar moieties on the sidechains has been reported (Rümenapf et al., 1993). Jordan et al. (2002a) demonstrated that inhibition of-glucosidase removal of terminal glucose moieties
from the nascent E2 glycoprotein prevented golgi processing of the E2 glycoprotein with the loss of production of infectious progeny virus.
RNA REPLICATION
Replication of the BVDV genomic RNA occurs downstream of polyprotein translation, being dependent on the presence of viral proteins that function specifically in RNA replication. Replication of the genomic RNA involves the copying of the infecting plus-sense RNA into minus-sense RNA copies, forming the replicative form (RF). The negative-strand of the RF is copied by stranddisplacement, giving rise to the RNA replicative intermediate (RI), and releasing the displaced plussense RNA strand when nascent strand transcription is complete. The RF is reutilized, primarily by recycling of the negative-sense. The kinetics of synthesis showed that the rate of synthesis of the positivesense strand at 12 hours postinfection was 10 times that of the negative strand (Gong et al., 1996). At any given time, there were approximately 6–7 nascent plus-stranded RNA strands on the template RI (Gong et al., 1998).
Viral proteins known to participate in RNA repli-